Highly Sensitive Microfilter Detects Trace Amounts of Circulating Tumor Cells in Whole Blood
Cancer cell count in the blood of cancer patients is extremely low. If these cells are easily detectable, cancer diagnosis may be possible by simply using a blood test, thus reducing patient burden.
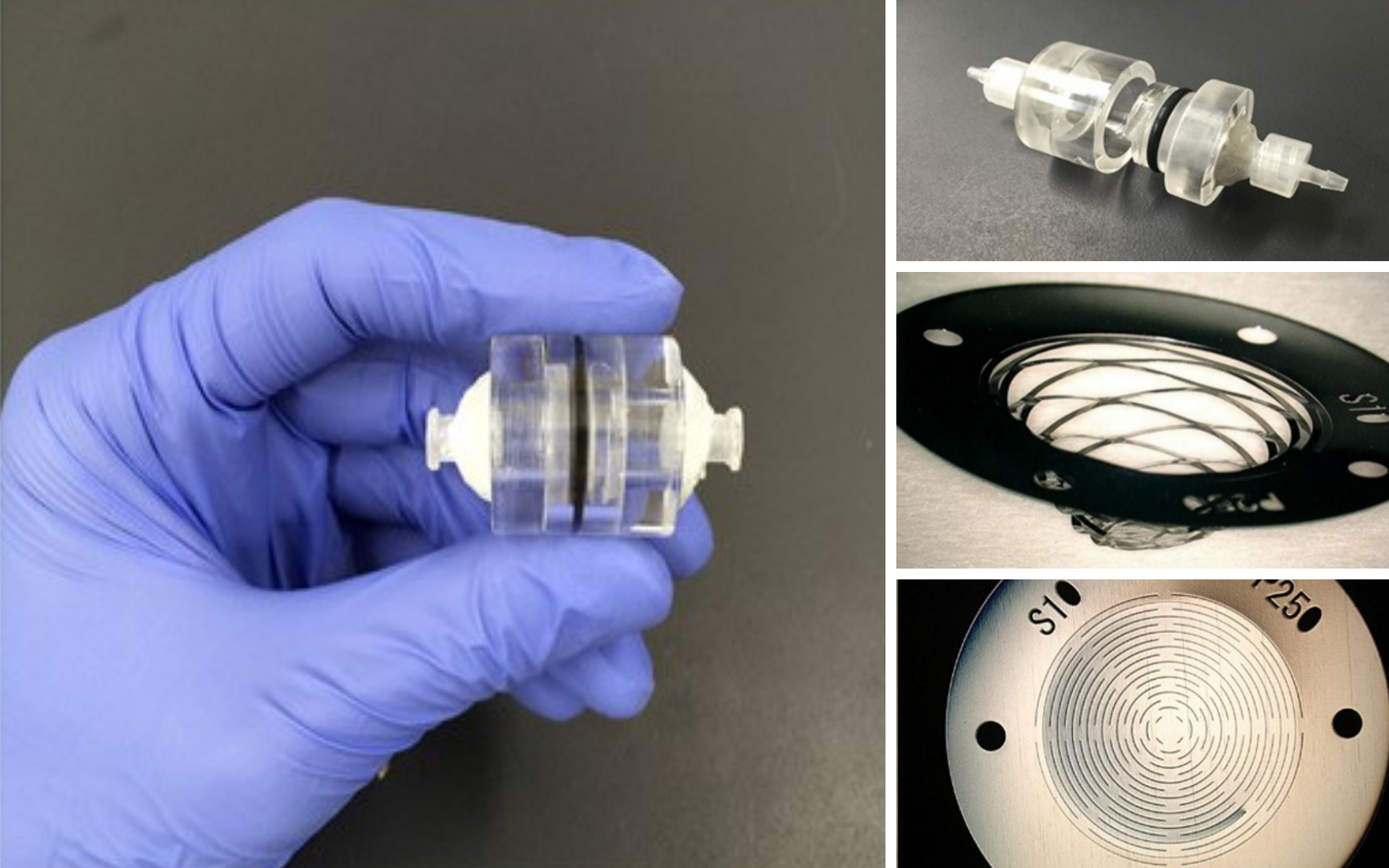
A microfilter device that can easily separate and capture trace amounts of cancer cells in blood has been developed by a Kumamoto University research group. The highly sensitive microfilter can successfully work in samples containing as few as five tumor cells in 1 mL of blood and does not require expensive equipment or reagents, unlike certain pre-existing cell capture technologies.
The palm sized device is expected to contribute to the development of new cancer diagnostic technologies based on cancer cells in the blood, such as early detection by a blood test, postoperative management, and recurrence monitoring.
The blood of people with cancer contains trace amounts of Circulating Tumor Cells (CTCs) that have detached from the primary cancer site and travel through the bloodstream.
However, the amount of these cells is only a few per milliliter, whereas red or white blood cells number in the billions, making it very difficult to separate and detect them.
Although devices for detecting Circulating Tumor Cells (CTCs) have been developed in the past, they require expensive equipment and reagents, which has been a bottleneck for their practical application.
If the cells can be successfully isolated from a patient’s blood, they can provide an early warning sign that cancer is present, as well as a minimally invasive diagnostic marker, requiring only a small blood sample.
However, isolating the cells is easier said than done, as they are typically only present in very low concentrations. There may only be several of the circulating tumor cells in a milliliter of blood, whereas the number of white and red blood cells could easily number in the billions. A needle in a haystack doesn’t even come close.
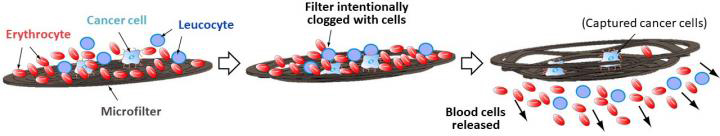
The unique microfilter device developed by the Kumamoto University research group can easily and inexpensively separate and capture Circulating Tumor Cells (CTCs) without any large equipment.
Dynamic Deformable Microfilter
The device is dynamically and three-dimensionally deformed by the fluid force when blood is pumped through it. It also utilizes nucleic acid aptamers, which bind specifically and firmly to target molecules. This enables both size-selective and affinity-selective separation and capture of tiny cancer cells.
While evaluating the device, researchers demonstrated that it could capture cancer cells even at a concentration of just five cancer cells in one mL of healthy blood. Since there are about five billion (red and white) blood cells in an mL of blood, the device proved to have a very high detection capability.

Researchers also found that almost no blood cells were adsorbed by the microfilter, achieving a blood cell removal rate of more than 98%. It also had a highly selective detection capability. Furthermore, in a comparative evaluation, they proved that the device was able to detect cancer cells with higher accuracy than existing devices.
When blood flows over the microfilter, cancer cells in the blood bind to the nucleic acid aptamer. The hydrodynamic force of the blood opens the slits of the microfilter and the blood cells flow away, leaving the cancer cells on the microfilter.
“This work demonstrates that our microfilter device can accurately detect trace amounts of cancer cells in the blood,” said Associate Professor Yuta Nakashima, who led the study.
“We expect it will be adopted for cancer diagnosis and treatment, including for early diagnosis of cancers that cannot be detected by imaging like CT and PET scans, postoperative follow-up, recurrence monitoring, and tailor-made treatments. In the future, we plan to use blood samples donated by cancer patients to verify the practical and clinical application of the method.”
Primary References
- Kumamoto University
- Detection of cancer cells in whole blood using a dynamic deformable microfilter and a nucleic acid aptamer. Fukuyama S, Kumamoto S, Nagano S, Hitotsuya S, Yasuda K, Kitamura Y, Nakashima Y. (2021).Talanta, 228, 122239. DOI: 10.1016/j.talanta.2021.122239